The dynamic behavior of microtubules (MTs) is essential to their functions and a large number of cellular factors regulate it during the cell cycle. We aim to define the structural basis of dynamic instability, the essential process that, fueled by GTP hydrolysis, leads to switching between MT growth and depolymerization, and that is inhibited by chemotherapeutics like taxol (Fig. 1). Our original studies of tubulin bound to taxol in polymerized, straight protofilaments, obtained by electron crystallography established the structural basis of nucleotide exchange and polymerization-coupled hydrolysis. Our lab later obtained two structures proposed to mimic intermediates in the assembly and disassembly of microtubules that illustrate the conformational consequences of the nucleotide state and how they relate to longitudinal and lateral assembly.
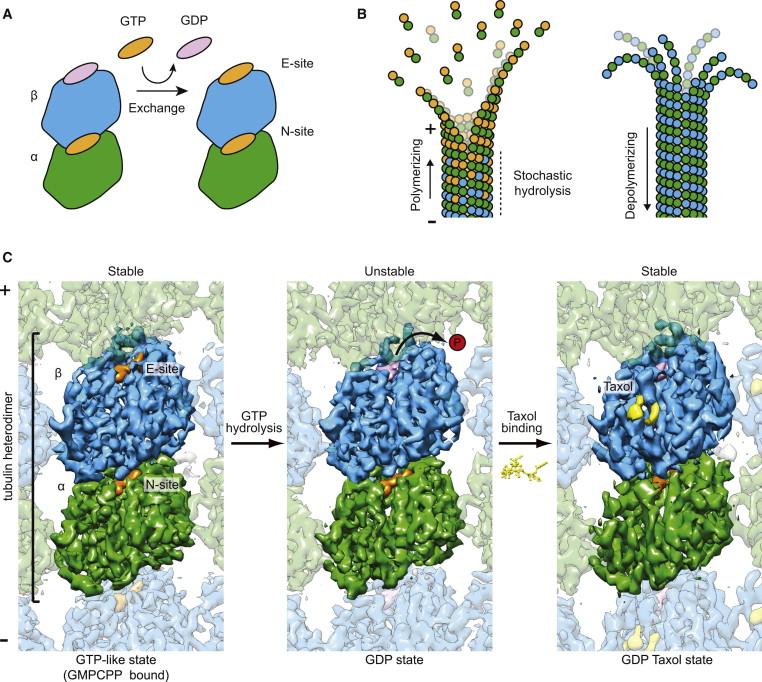
Later on our studies have centered on defining the conformational changes within the microtubule upon GTP hydrolysis. Our studies showed that GTP hydrolysis results in a compaction at the interdimer longitudinal interface that buries the E-site nucleotide and a conformational change in alpha tubulin that generates strain in the MT lattice. More recently, we have obtained six cryo-EM structures at ~3.5 Å resolution of MTs bound to GMPCPP, GTPγS, or GDP, either decorated with kinesin or copolymerized with the +TIP protein EB3. This resolution has allowed us to derive atomic models directly from the density maps and to shed unique light on several key aspects of MT structure and dynamics
Our studies have shown that through the GTP hydrolysis cycle, subtle changes around the E-site nucleotide trigger conformational changes in in accumulation site. The structures also revealed, for the first time, the atomic details of the native lateral contacts between protofilaments, and showed that they do not change with GTP hydrolysis. Reconstructions without any symmetry applied (and reaching 3.9-4.4 Å resolution), allowed us the detailed visualization of the seam naturally present in the MT lattice. We found that the relative position of the two PFs at the seam makes this region of the MT deviate from the cylindrical geometry observed for the rest of the tube. This deviation is more significant for less stable MTs (i.e. GDP versus GMPCPP), suggesting that the seam may be a weak point in the MT lattice that could play an important role in MT disassembly.
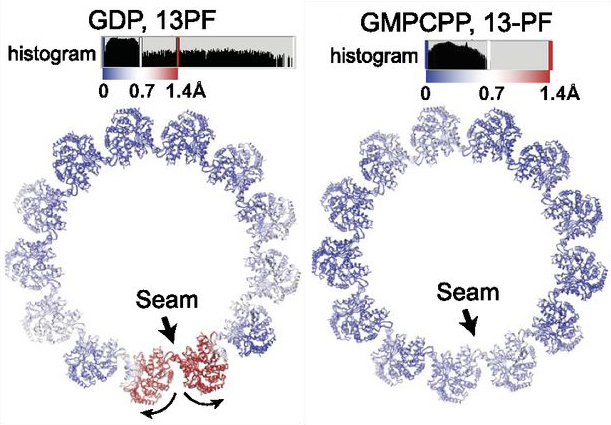
After our original studies using different GTP analogs and MTs bound to either kinesin or EB3[5, 6], which showed that hydrolysis gives rise to a compaction of the dimer and a change in the skew angle of PFs, we developed a method that allowed structure determination of MTs without decoration by other proteins (to differentiate and -tubulin)[29]. Our structures of MTs bound to GMPCPP, GDP, and GTPγS in the absence of binding proteins allowed us to differentiate the effects of nucleotide state versus MAP binding on MT structure. We also found that in the GDP-MT, PFs separate at the seam, while this is not the case for GMPCCP and GTPgS-stabilized MTs, suggesting that an “opening” of the seam after hydrolysis likely contributes to MT destabilization. In our model, the MT lattice integrates internal (i.e., nucleotide state) and outside signals (i.e., binding of MAPs, forces) resulting in rearrangements that in turn affect the affinity of other MT partners and result in the exquisite regulation of MT dynamics.
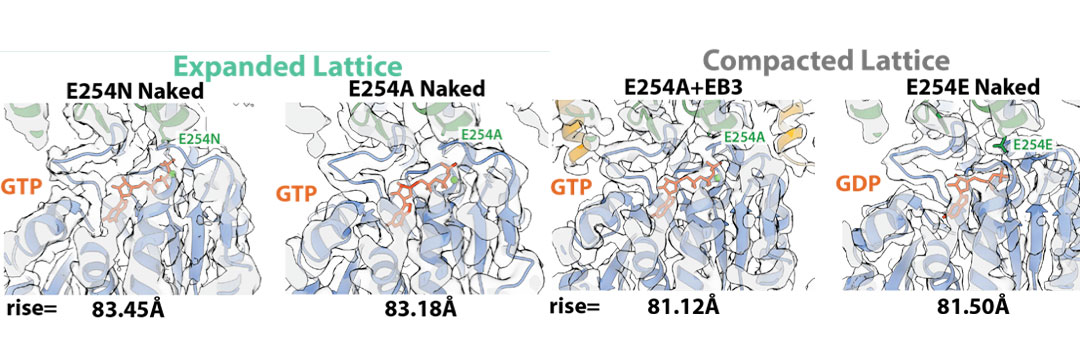
Structure of MTs from hydrolysis mutant tubulin – Given that different non-hydrolyzable analogs (i.e. GMPCPP and GTPγS) give rise to different tubulin/MT structures, we collaborated with Thomas Surrey (CRG, Barcelona) to generate a GTP-bound MT lattice using recombinant human tubulin mutated to prevent GTP hydrolysis. We found that two different catalytically dead mutants (E254A, and E254N) have an expanded lattice (left), similar to that previously observed for porcine GMPCPP-MTs. We also found that EB proteins can compact these GTP-bound MTs (right). The seam appears to be stabilized when bound to GTP and destabilized when bound to GDP, further supporting the idea of its involvement in the depolymerization process.
Currently we are further investigating how cellular factors and in particular microtubule plus end binding proteins regulate dynamic instability.
See also:
Eva Nogales, Sharon G. Wolf, Kenneth H. Downing (1998). Structure of the αβ-tubulin dimer by electron crystallography. Nature. 391, 199-203.
Eva Nogales, Kenneth H. Downing, Linda A Amos, Jan Löwe. (1998) Tubulin and FtsZ form a distinct family of GTPases. Nat Struct Biol. 6, 451-8.
Eva Nogales, M Whittaker, RA Milligan, Kenneth H. Downing. (1999) High-resolution model of the microtubule. Cell. 96(1), 79-88.
- Löwe, H. Li, K.H. Downing, E. Nogales. (2001) Refined Structure of αβ-Tubulin at 3.5 Å Resolution.J Mol Biol.313, 1045-1057.
Hong-Wei Wang and Eva Nogales. (2005) The nucleotide-dependent bending flexibility of tubulin regulates microtubule assembly. Nature. 435, 911-915.
Gregory M. Alushin, Gabriel C. Lander, Elizabeth H. Kellogg, Rui Zhang, David Baker, and Eva Nogales (2014) High resolution microtubule structures reveal the structural transitions in αβ–tubulin upon GTP hydrolysis Cell, 157(5)1117-29.
Nogales, E. (2015) An Electron Microscopy Journey in the Study of Microtubule Structure and Dynamics. Protein Sci. 24, 1912-1919.
Zhang, R., Alushin, G.M., Brown, A. and Nogales E. (2015) Mechanistic origin of microtubule dynamic instability and its regulation by EB proteins. Cell. 162, 849-859.
Kellogg, E., Howes, S., Ti, S-C., Ramirez-Aportela, E., Kapoor, T., Chacon, P. and Nogales, E. (2016) Near-atomic resolution cryo-EM structure of PRC1 bound to the microtubule. PNAS 113, 9430-9439.
Nogales, E. (2016) Dear microtubule, I see you. Mol. Bol. Cell 27 , 3202-3204.
Borisy, G, Heald, R., Howard J., Janke, C Musacchio, A. and Nogales, E. (2016) Microtubules: 50 years on from the discovery of tubulin. Nat. Rev. Mol. Cell Biol. 17, 322-328.
Nogales E. and Zhang, R. (2016) Visualizing microtubule structure and interactions. Curr. Opi. Struct. Biol. 37, 90-96.
Howes, S.C., Geyer, E.A., LaFrance, B., Zhang, R., Kellogg, E.H., Westermann, S., Rice, L.M. and Nogales, E. (2017) Structural differences between yeast and mammalian microtubules revealed by cryo-EM. JCB 216 , 2669-2677.
Kellogg, E., Hejab, N.M.A., Howes, S., Northcote, P, Miller, J.H., Diaz, J.F., Downing, K.H. and Nogales, E. (2017). Insights into the distinct mechanisms of action of taxane and non-taxane microtubule stabilizers from cryo-EM studies. J. Mol. Biol. 429, 633–646. Cover in that issue.
Zhang, R., LaFrance, B. & Nogales, E. (2018) Separating the effects of nucleotide and EB binding on microtubule structure. Proc. Natl. Acad. Sci. U.S.A. 115, E6191–E6200.
Eshun-Wilson, L., Zhang, R., Portran, D., Nachury, M. V., Toso, D. B., Löhr, T., Vendruscolo, M., Bonomi, M., Fraser, J. S. & Nogales, E (2019). Effects of α-tubulin acetylation on microtubule structure and stability. Proc. Natl. Acad. Sci. U.S.A. 116, 10366–10371.
LaFrance, B. J., Roostalu, J., Henkin, G., Greber, B. J., Zhang, R., Normanno, D., McCollum, C. O., Surrey, T. & Nogales, E. (2022) Structural transitions in the GTP cap visualized by cryo-electron microscopy of catalytically inactive microtubules. Proc. Natl. Acad. Sci. U.S.A. 119.
