Microtubules (MTs) are highly conserved cytoskeletal polymers that are essential for cell growth, cell division, and intracellular transport. (MT) Microtubule-associated proteins, or MAPs, are architects of specialized MT populations for diverse functions in the cell.
Based on their mode of action, MAPs can be grouped into the following groups: (1) motile MAPs, or motor proteins, such as kinesin and dynein, that generate forces and movement; (2) enzymes that break or depolymerize microtubules; (3) microtubule nucleators (4) end-binding proteins that specifically associate with +/- ends of microtubules; and (5) “structural” MAPs. “Structural” MAPs are generally defined as proteins that bind and stabilize MTs, although a clear, systemic view of their functions is missing. Initially, structural MAPs were defined as proteins that: co-polymerize with tubulin through repeated cycles of MT assembly in vitro; do not associate with any other brain subcellular fraction other than microtubules; stain the same fibrous elements in cultured cells as anti-tubulin in double-label immunofluorescence experiments; form a dispersed staining pattern when cytoplasmic MTs are disrupted by colchicine. Yet, when over expressed, some proposed MAPs may be coerced into a non-physiological MT interaction, so additional biochemical assays are necessary to assess the activity of a structural MAPs.
Structural MAPs can directly shape the cytoskeleton in ways such as promoting MT bundling, cross-linking multiple cytoskeletal filaments, or stabilizing and polymerizing MTs.
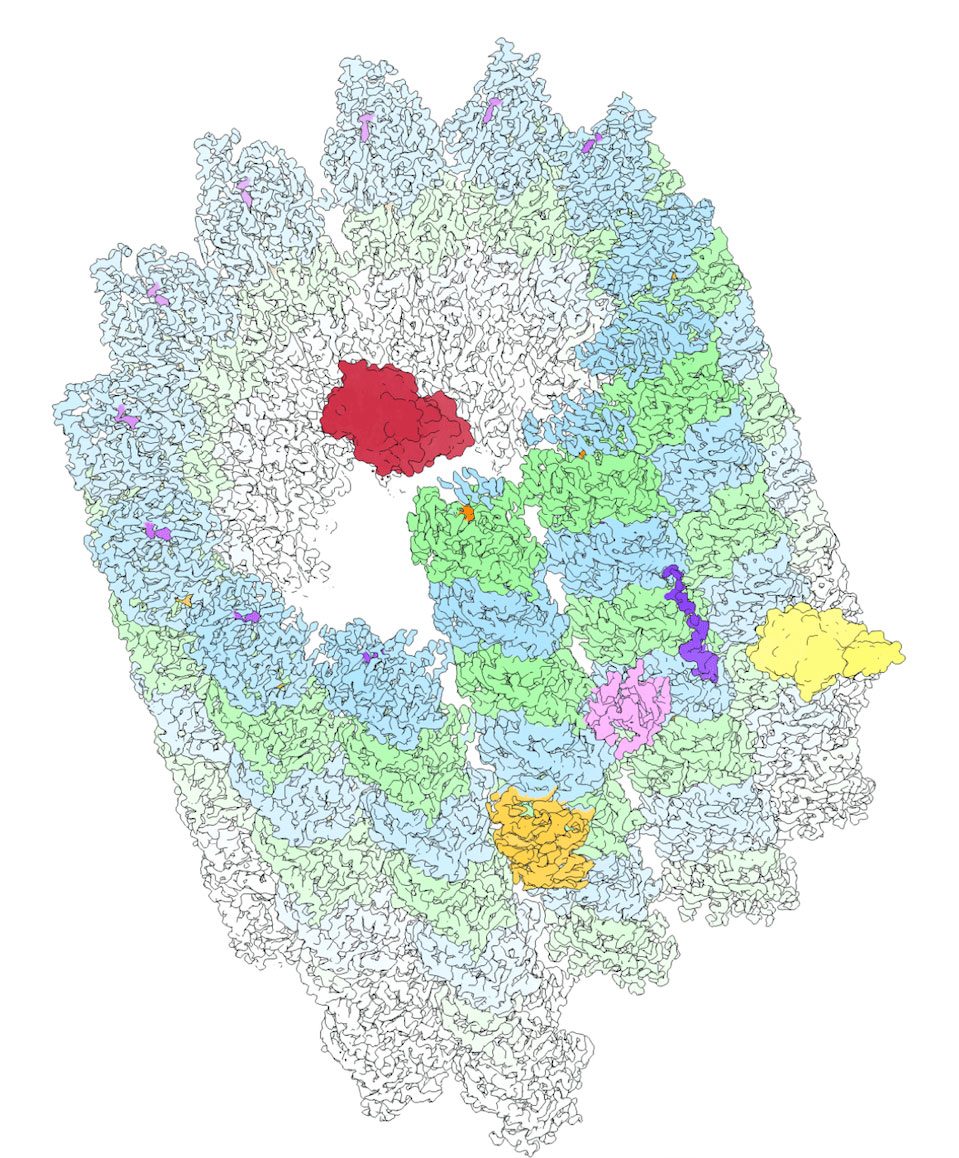
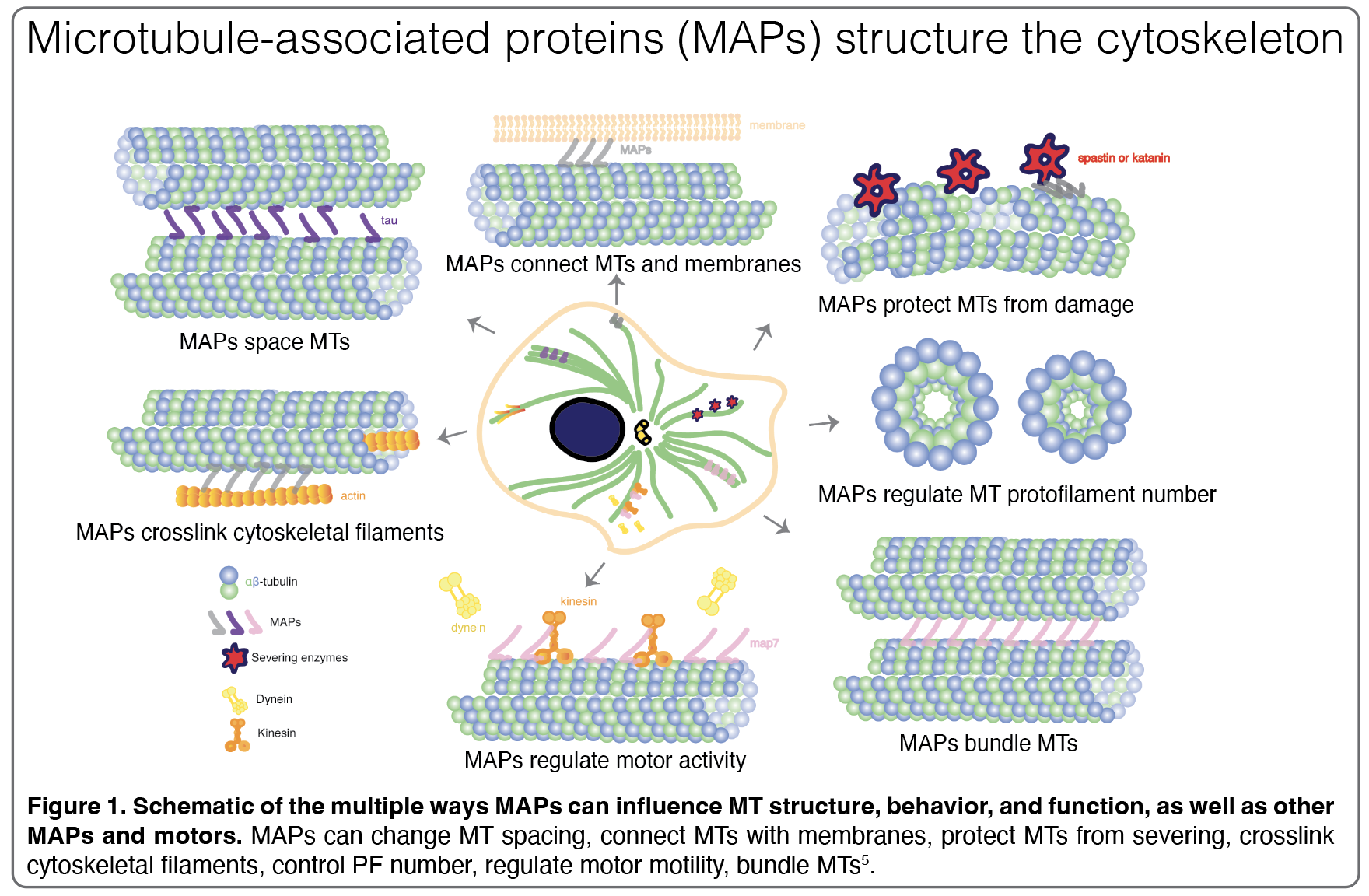
They share a positively charged MT-binding domain (MTBD) and projection domain and not only influence MT structure and behavior but may also drive interactions with other MAPs. Since their discovery in the 1970s, multiple hypotheses have formed to describe how structural MAPs can anchor and manipulate MT populations. First, MAPs could physically link neighboring MTs or, as a second hypothesis, MAP stabilization of MTs could be enough to trigger bundling, for instance, by shielding the negatively charged C-terminal tails, reducing electrostatic repulsion between MTs, and therefore promoting spontaneous MT bundling. When the C-terminal tails are cleaved from the MT surface using subtilisin, thus neutralizing the negative surface charge along the MT lattice, scientists see that MTs bundle immediately, which provides strong evidence for the latter hypothesis.
Multiple reports show MTBDs of MAPs alone are sufficient to neutralize the C-terminal tails, so the projection domain may play a prominent role in dictating the primary function of the MAP. For example, the MTBD of MAP4 alone induces MT bundling but full-length MAP4, which has a long projection domain, binds to MTs but does not induce MT bundling. Interestingly, proteins with similar structures may not bind the same site on the MT: the CH domain of the end-binding proteins binds between protofilaments, while the CH domain of the Ndc80 protein, a component of the kinetochore complex, binds along protofilaments. These complexities highlight the hidden conformational heterogeneity and structural plasticity of the MT lattice. Moreover, earlier in vivo studies showed that MT spacing is regulated by projection domain size and additional studies found that certain MAPs express isoforms with projection domains that vary in length, suggesting that MAPs may use their projection domains to perform unique functions in the cell. However, the mechanism of how projection domains drive MT-MAP and MAP-MAP interactions remains elusive.
The structural MAPs tau and MAP7
MAPs can influence MT function and share a positively charged MT-binding domain (MTBD) and floppy projection domain, which can range in size. Tau stands out as one of the most abundant MAPs, constituting nearly 80% of all neuronal MAPs. Tau stabilizes and bundles MTs and is developmentally regulated. Full-length adult tau is intrinsically disordered and contains a N-terminal projection domain, a MTBD, two proline-rich regions (120-197; 197-242), and four variable sequence repeats (R1 to R4). Multiple life-threating neurodegenerative diseases, including Alzheimer’s, develop when misregulated tau loss affinity for MTs and creates filamentous aggregates called neurofibrillary tangles. Early studies of tau predicted that it interacted with MTs in an intrinsically disordered manner, while others proposed it adopted a helical structure upon MT-binding. Other studies showed that tau bound along PFs when added to pre-formed MTs as well as, surprisingly, inside the lumen of MTs near the Taxol-binding site when co-polymerized with MTs.
In 2018, our group successfully resolved the structure of tau to ~3 Å and showed that it did, in fact, interact with MTs and stabilize outer, longitudinal MT contacts. Structures of individual tau repeats in complex with MTs lead to a model in which each tau repeat extends across both intra- and interdimer interfaces, centered on α-tubulin and connecting three tubulin monomers. Kellogg et al. 2018 found that each repeat tau has two MT-binding sites, with the major site residing on the “anchor point”, an area on tubulin that remains unchanged during the structural transitions associated with nucleotide hydrolysis or MT assembly and disassembly, which explains how tau can form tubulin rings and copurify with MTs through tubulin assembly-disassembly cycles. Moreover, with each addition of the tandem repeats, scientists saw tau’s affinity for the MT lattice increase.
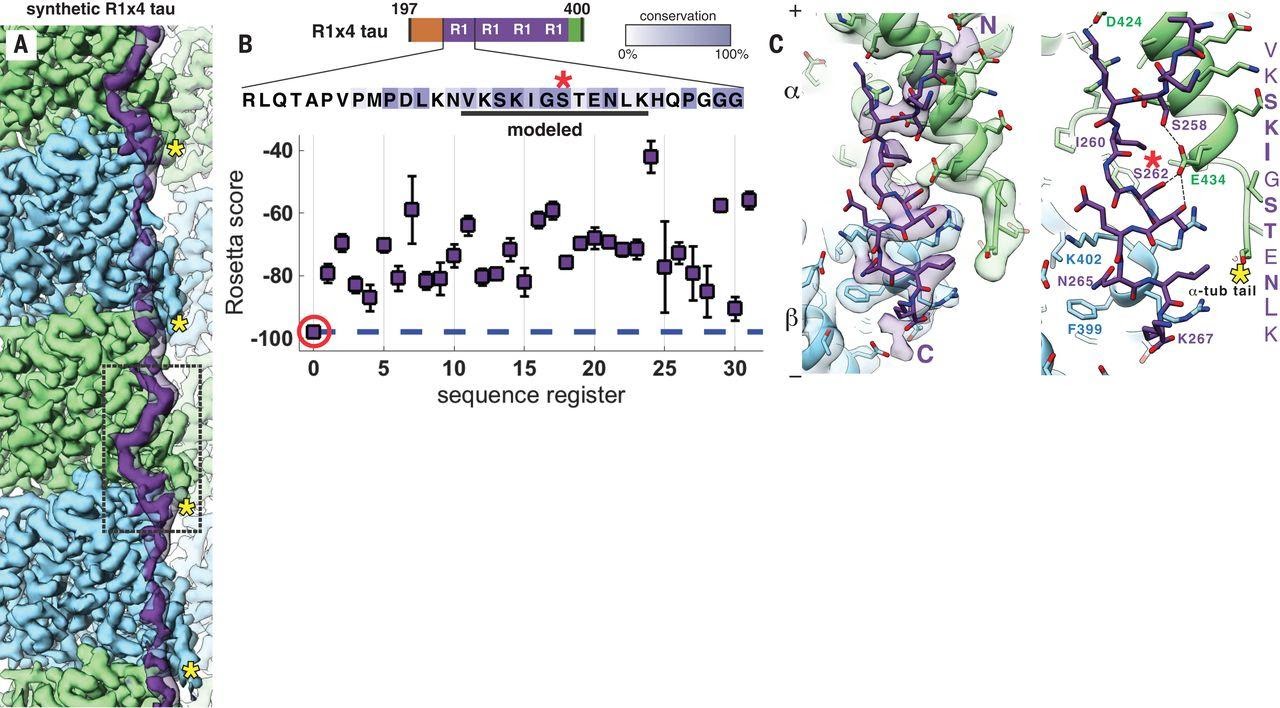
Thus, cells can regulate both the number of repeats and the length of the projection domain to modulate tau’s affinity for the MT lattice. The tandem binding of tau repeats along PFs also elucidates how tau stabilizes and promotes MT polymerization. Specifically, tau tethers multiple tubulin dimers together across longitudinal interfaces, which provides insight into the reduced off-rate of tubulin dimers in the presence of tau, therefore, reduced MT catastrophe events. Though, an open question in the field, is how other structural elements of tau, such as its projection domain, contribute to MT stability, inter-MT interactions, and other MAP interactions. Kellogg et al. 2018 also superimposed tau with kinesin to show that tau’s proposed inhibitory effect on kinesin may be caused by binding overlap, but this interaction has yet to be fully characterized. Investigating the projection domain of other MAPs and/or how other MAPs or motors engage with tau’s projection domain may be a means to unlock how tau’s projection domain may confer MT stability.
Interestingly, MAP7 was shown to not only compete with other maps, like tau, but also with molecular motors – highlighting a novel functional duality of this MAP. MAP7 is largely intrinsically disordered and contains two positively charged MTBDs, MTDB and p123, a proline rich region, a C-terminal projection domain that encompasses a kinesin-binding domain, and two projected coiled-coil domains with one in the MTDB and the second in the kinesin-binding domain. Originally discovered in HeLa cells, MAP7 is widely conserved in a range of organisms, from Drosophila to mice. In Drosophila, MAP7, or Enconsin/EMAP-115, has been shown to regulate cell polarity of oocytes, nuclear positioning in muscle cells, organelle transport in S2 cells, and spindle morphogenesis in neural stem cells. Moreover, in mice, MAP7 has also been found to regulate axon collateral branch development in sensory neurons in rodents. Intriguingly, MAP7 and tau compete to bind the MT lattice yet have opposite effects on kinesin: MAP7 enhances kinesin binding while tau inhibits it. A recent study proposed that MAP competition may be a mechanism the cell exploits to coordinate motor activity, the MT cytoskeleton and, ultimately, cell fate.
Excitingly, in collaboration with the Yildez lab, our lab set out to uncover the molecular mechanism of MAP7-mediated motor regulation. Lab members Qianglin Fang and Lisa Eshun-Wilson worked together to determine the first near-atomic structure of MAP7 on the microtubule, now published in Science (2022).
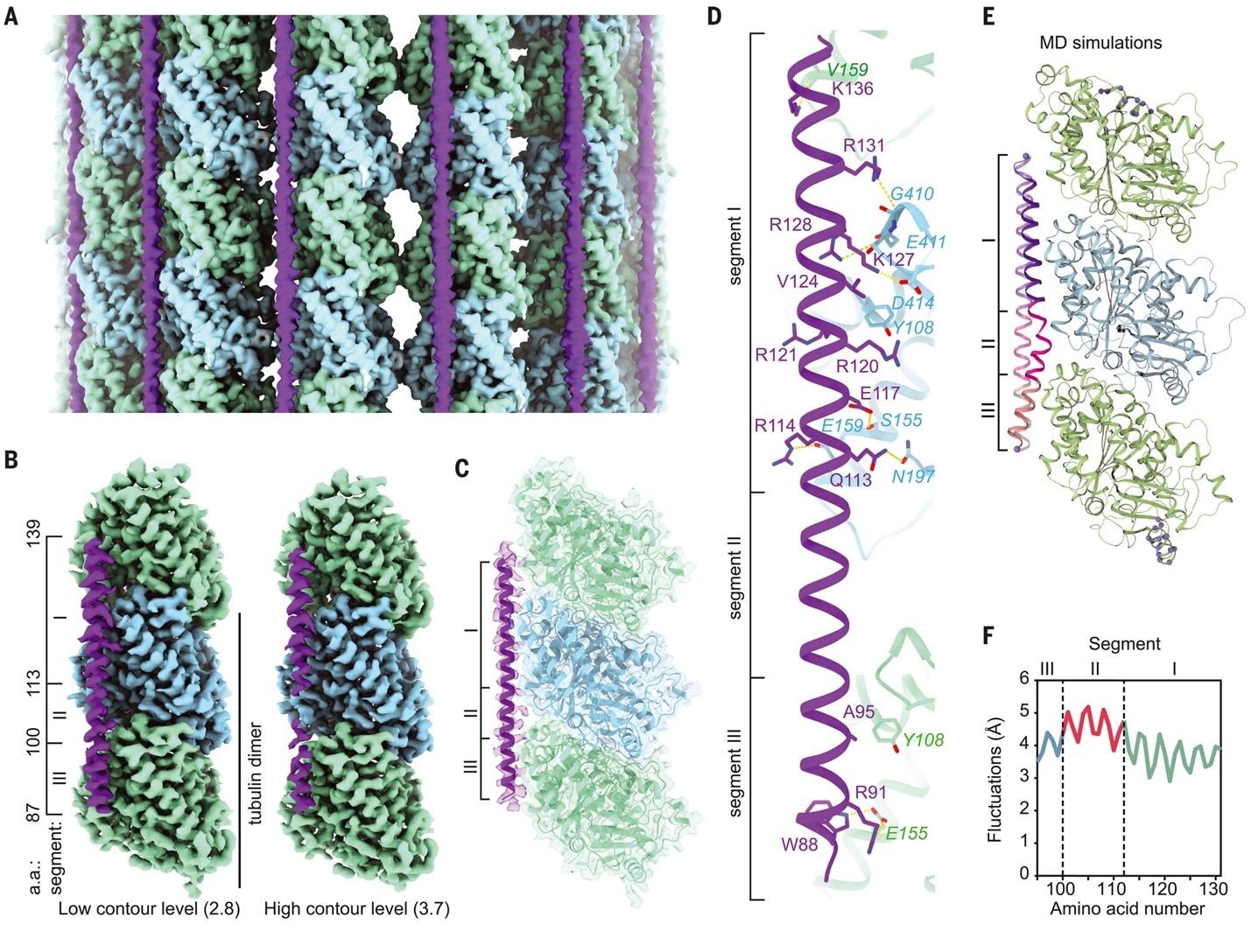
They found that the microtubule-binding site of MAP7 overlaps with kinesin. However, by tethering kinesin-1 to the microtubule, the MAP7 projection domain enabled the motor to diffuse to available sites on microtubules partially decorated by MAP7. Based on these findings, we propose a model for kinesin stepping on MAP7-decorated MTs. MAP7 recruits kinesin-1 to the MT and activates subsequent motility. MT binding of MAP7 also obstructs kinesin stepping along the protofilament, which results in kinesin dissociation from the MT. However, the MAP7 projection domain tethers kinesin to the MT surface, allowing it to rebind the MT at nearby sites not blocked by its MTBD. This “tethered diffusion” of kinesin appears as large forward, sideways, or backward displacements in our trajectories.
